Pre-trained ODAC models are versatile across various MOF-related tasks. To begin, we’ll start with a fundamental application: calculating the adsorption energy for a single CO2 molecule. This serves as an excellent and simple demonstration of what you can achieve with these datasets and models.
For predicting the adsorption energy of a single CO2 molecule within a MOF structure, the adsorption energy () is defined as:
Each term on the right-hand side represents the energy of the relaxed state of the indicated chemical system. For a comprehensive understanding of our methodology for computing these adsorption energies, please refer to our paper.
Loading Pre-trained Models¶
Need to install fairchem-core or get UMA access or getting permissions/401 errors?
Install the necessary packages using pip, uv etc
! pip install fairchem-core fairchem-data-oc fairchem-applications-cattsunamiGet access to any necessary huggingface gated models
Get and login to your Huggingface account
Request access to https://
huggingface .co /facebook /UMA Create a Huggingface token at https://
huggingface .co /settings /tokens/ with the permission “Permissions: Read access to contents of all public gated repos you can access” Add the token as an environment variable using
huggingface-cli loginor by setting the HF_TOKEN environment variable.
# Login using the huggingface-cli utility
! huggingface-cli login
# alternatively,
import os
os.environ['HF_TOKEN'] = 'MY_TOKEN'A pre-trained model can be loaded using FAIRChemCalculator. In this example, we’ll employ UMA to determine the CO2 adsorption energies.
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p2")
calc = FAIRChemCalculator(predictor, task_name="odac")WARNING:root:device was not explicitly set, using device='cuda'.
Adsorption in rigid MOFs: CO2 Adsorption Energy in Mg-MOF-74¶
Let’s apply our knowledge to Mg-MOF-74, a widely studied MOF known for its excellent CO2 adsorption properties. Its structure comprises magnesium atomic complexes connected by a carboxylated and oxidized benzene ring, serving as an organic linker. Previous studies consistently report the CO2 adsorption energy for Mg-MOF-74 to be around -0.40 eV [1] [2] [3].
Our goal is to verify if we can achieve a similar value by performing a simple single-point calculation using UMA. In the ODAC23 dataset, all MOF structures are identified by their CSD (Cambridge Structural Database) code. For Mg-MOF-74, this code is OPAGIX. We’ve extracted a specific OPAGIX+CO2 configuration from the dataset, which exhibits the lowest adsorption energy among its counterparts.
import matplotlib.pyplot as plt
from ase.io import read
from ase.visualize.plot import plot_atoms
mof_co2 = read("structures/OPAGIX_w_CO2.cif")
mof = read("structures/OPAGIX.cif")
co2 = read("structures/co2.xyz")
fig, ax = plt.subplots(figsize=(5, 4.5), dpi=250)
plot_atoms(mof_co2, ax)
ax.set_axis_off()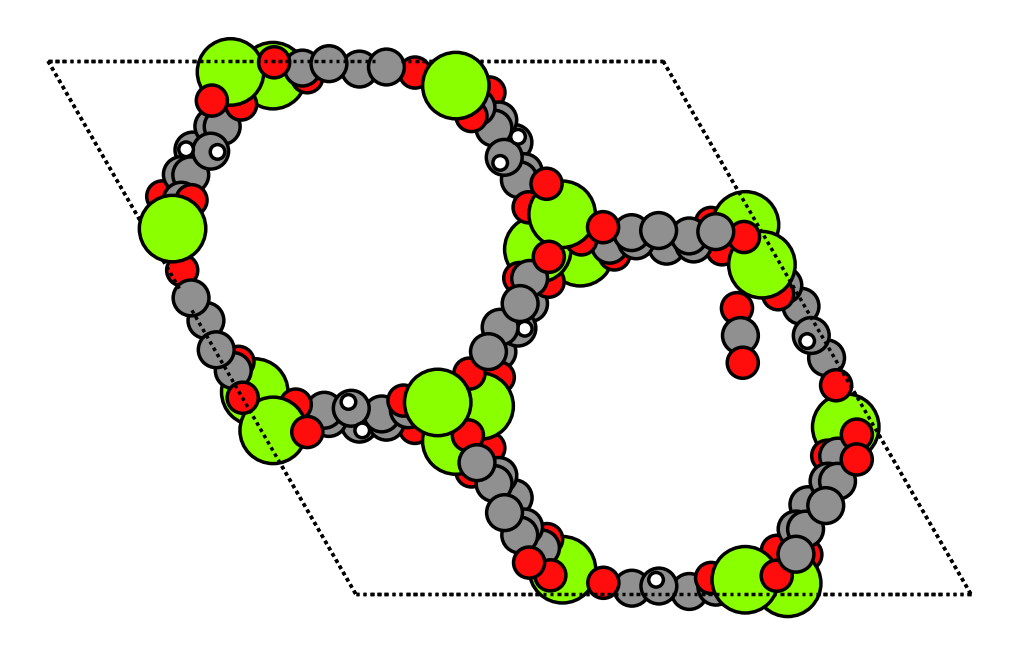
The final step in calculating the adsorption energy involves connecting the FAIRChemCalculator to each relaxed structure: OPAGIX+CO2, OPAGIX, and CO2. The structures used here are already relaxed from ODAC23. For simplicity, we assume here that further relaxations can be neglected. We will show how to go beyond this assumption in the next section.
mof_co2.calc = calc
mof.calc = calc
co2.calc = calc
E_ads = (
mof_co2.get_potential_energy()
- mof.get_potential_energy()
- co2.get_potential_energy()
)
print(f"Adsorption energy of CO2 in Mg-MOF-74: {E_ads:.3f} eV")Adsorption energy of CO2 in Mg-MOF-74: -0.473 eV
Adsorption in flexible MOFs¶
The adsorption energy calculation method outlined above is typically performed with rigid MOFs for simplicity. Both experimental and modeling literature have shown, however, that MOF flexibility can be important in accurately capturing the underlying chemistry of adsorption [1] [2] [3]. In particular, uptake can be improved by treating MOFs as flexible. Two types of MOF flexibility can be considered: intrinsic flexibility and deformation induced by guest molecules. In the Open DAC Project, we consider the latter MOF deformation by allowing the atomic positions of the MOF to relax during geometry optimization [4]. The addition of additional degrees of freedoms can complicate the computation of the adsorption energy and necessitates an extra step in the calculation procedure.
The figure below shows water adsorption in the MOF with CSD code WOBHEB with added defects (WOBHEB_0.11_0) from a DFT simulation. A typical adsorption energy calculation would only seek to capture the effects shaded in purple, which include both chemisorption and non-bonded interactions between the host and guest molecule. When allowing the MOF to relax, however, the adsorption energy also includes the energetic effect of the MOF deformation highlighted in green.
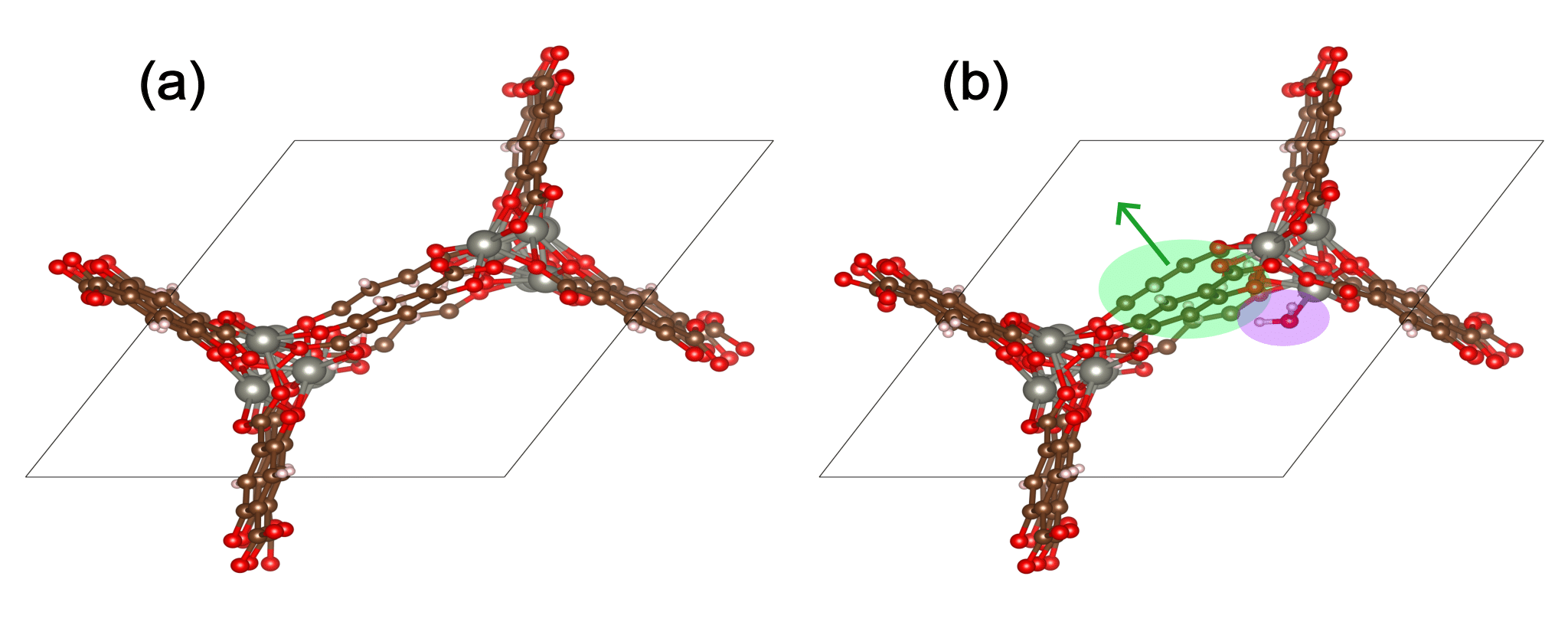
To account for this deformation, it is vital to use the most energetically favorable MOF geometry for the empty MOF term in Eqn. 1. Including MOF atomic coordinates as degrees of freedom can result in three possible outcomes:
The MOF does not deform, so the energies of the relaxed empty MOF and the MOF in the adsorbed state are the same
The MOF deforms to a less energetically favorable geometry than its ground state
The MOF locates a new energetically favorable geoemtry relative to the empty MOF relaxation
The first outcome requires no additional computation because the MOF rigidity assumption is valid. The second outcome represents physical and reversible deformation where the MOF returns to its empty ground state upon removal of the guest molecule. The third outcome is often the result of the guest molecule breaking local symmetry. We also found cases in ODAC in which both outcomes 2 and 3 occur within the same MOF.
To ensure the most energetically favorable empty MOF geometry is found, an addition empty MOF relaxation should be performed after MOF + adsorbate relaxation. The guest molecule should be removed, and the MOF should be relaxed starting from its geometry in the adsorbed state. If all deformation is reversible, the MOF will return to its original empty geometry. Otherwise, the lowest energy (most favorable) MOF geometry should be taken as the reference energy, , in Eqn. 1.
H2O Adsorption Energy in Flexible WOBHEB with UMA¶
The first part of this tutorial demonstrates how to perform a single point adsorption energy calculation using UMA. To treat MOFs as flexible, we perform all calculations on geometries determined by geometry optimization. The following example corresponds to the figure shown above (H2O adsorption in WOBHEB_0.11_0).
In this tutorial, corresponds to the energy of determined from geometry optimization of .
First, we obtain the energy of the empty MOF from relaxation of only the MOF:
import ase.io
from ase.optimize import BFGS
mof = ase.io.read("structures/WOBHEB_0.11.cif")
mof.calc = calc
relax = BFGS(mof)
relax.run(fmax=0.05)
E_mof_empty = mof.get_potential_energy()
print(f"Energy of empty MOF: {E_mof_empty:.3f} eV") Step Time Energy fmax
BFGS: 0 14:18:58 -1077.368917 0.129115
BFGS: 1 14:18:58 -1077.370393 0.075188
BFGS: 2 14:18:59 -1077.372342 0.145320
BFGS: 3 14:19:01 -1077.374555 0.111799
BFGS: 4 14:19:03 -1077.376094 0.074296
BFGS: 5 14:19:07 -1077.377454 0.063781
BFGS: 6 14:19:09 -1077.378942 0.080760
BFGS: 7 14:19:10 -1077.380756 0.096894
BFGS: 8 14:19:10 -1077.382638 0.078412
BFGS: 9 14:19:13 -1077.384447 0.086865
BFGS: 10 14:19:14 -1077.386286 0.083326
BFGS: 11 14:19:14 -1077.388395 0.084011
BFGS: 12 14:19:14 -1077.390738 0.069085
BFGS: 13 14:19:15 -1077.393129 0.076037
BFGS: 14 14:19:16 -1077.395559 0.084308
BFGS: 15 14:19:16 -1077.398149 0.079981
BFGS: 16 14:19:17 -1077.400825 0.079904
BFGS: 17 14:19:19 -1077.403368 0.067387
BFGS: 18 14:19:19 -1077.405674 0.070439
BFGS: 19 14:19:20 -1077.407933 0.087898
BFGS: 20 14:19:24 -1077.410401 0.083994
BFGS: 21 14:19:27 -1077.413125 0.059591
BFGS: 22 14:19:30 -1077.415977 0.071931
BFGS: 23 14:19:33 -1077.418823 0.067819
BFGS: 24 14:19:34 -1077.421564 0.069902
BFGS: 25 14:19:37 -1077.424156 0.067341
BFGS: 26 14:19:39 -1077.426518 0.060871
BFGS: 27 14:19:40 -1077.428604 0.069311
BFGS: 28 14:19:43 -1077.430414 0.060335
BFGS: 29 14:19:44 -1077.431999 0.051494
BFGS: 30 14:19:46 -1077.433387 0.056303
BFGS: 31 14:19:48 -1077.434618 0.057606
BFGS: 32 14:19:49 -1077.435738 0.046098
Energy of empty MOF: -1077.436 eV
Next, we add the H2O guest molecule and relax the MOF + adsorbate to obtain .
mof_h2o = ase.io.read("structures/WOBHEB_H2O.cif")
mof_h2o.calc = calc
relax = BFGS(mof_h2o)
relax.run(fmax=0.05)
E_combo = mof_h2o.get_potential_energy()
print(f"Energy of MOF + H2O: {E_combo:.3f} eV") Step Time Energy fmax
BFGS: 0 14:19:49 -1091.661287 1.120236
BFGS: 1 14:19:50 -1091.679632 0.313939
BFGS: 2 14:19:52 -1091.683944 0.232091
BFGS: 3 14:19:55 -1091.695506 0.302363
BFGS: 4 14:19:58 -1091.701044 0.210337
BFGS: 5 14:20:00 -1091.707224 0.171323
BFGS: 6 14:20:01 -1091.712981 0.183105
BFGS: 7 14:20:01 -1091.720514 0.262596
BFGS: 8 14:20:02 -1091.727865 0.202898
BFGS: 9 14:20:02 -1091.735396 0.175141
BFGS: 10 14:20:03 -1091.743448 0.214445
BFGS: 11 14:20:07 -1091.752660 0.253309
BFGS: 12 14:20:08 -1091.762639 0.232780
BFGS: 13 14:20:09 -1091.773121 0.197356
BFGS: 14 14:20:10 -1091.784458 0.164080
BFGS: 15 14:20:11 -1091.796075 0.252751
BFGS: 16 14:20:12 -1091.806469 0.270299
BFGS: 17 14:20:13 -1091.815242 0.186062
BFGS: 18 14:20:14 -1091.822969 0.130834
BFGS: 19 14:20:18 -1091.830275 0.120371
BFGS: 20 14:20:20 -1091.837494 0.140491
BFGS: 21 14:20:20 -1091.844736 0.154595
BFGS: 22 14:20:21 -1091.851973 0.162196
BFGS: 23 14:20:21 -1091.858841 0.163076
BFGS: 24 14:20:22 -1091.864336 0.156147
BFGS: 25 14:20:23 -1091.868622 0.434226
BFGS: 26 14:20:27 -1091.873920 0.216139
BFGS: 27 14:20:28 -1091.879918 0.103492
BFGS: 28 14:20:28 -1091.884370 0.089738
BFGS: 29 14:20:29 -1091.888782 0.149156
BFGS: 30 14:20:35 -1091.893305 0.142163
BFGS: 31 14:20:36 -1091.899423 0.199021
BFGS: 32 14:20:36 -1091.905042 0.237880
BFGS: 33 14:20:37 -1091.908028 0.462781
BFGS: 34 14:20:38 -1091.913662 0.159976
BFGS: 35 14:20:39 -1091.919533 0.098934
BFGS: 36 14:20:41 -1091.926837 0.131946
BFGS: 37 14:20:41 -1091.933437 0.365912
BFGS: 38 14:20:42 -1091.938593 0.218758
BFGS: 39 14:20:42 -1091.945577 0.347225
BFGS: 40 14:20:46 -1091.952173 0.190654
BFGS: 41 14:20:47 -1091.957377 0.498817
BFGS: 42 14:20:49 -1091.965799 0.278090
BFGS: 43 14:20:52 -1091.978220 0.274013
BFGS: 44 14:20:56 -1091.988356 0.203465
BFGS: 45 14:20:59 -1092.000088 0.626668
BFGS: 46 14:20:59 -1092.006514 0.763307
BFGS: 47 14:21:00 -1092.025725 0.284805
BFGS: 48 14:21:00 -1092.043554 0.395779
BFGS: 49 14:21:01 -1092.062259 0.882020
BFGS: 50 14:21:02 -1092.086192 0.414512
BFGS: 51 14:21:03 -1092.111293 0.291057
BFGS: 52 14:21:06 -1092.129334 0.732854
BFGS: 53 14:21:06 -1092.143992 0.430138
BFGS: 54 14:21:07 -1092.164359 0.374092
BFGS: 55 14:21:08 -1092.177481 0.325447
BFGS: 56 14:21:09 -1092.193323 0.228512
BFGS: 57 14:21:12 -1092.207749 0.251896
BFGS: 58 14:21:13 -1092.221841 0.333958
BFGS: 59 14:21:16 -1092.234015 0.399857
BFGS: 60 14:21:17 -1092.245236 0.390004
BFGS: 61 14:21:17 -1092.256708 0.282643
BFGS: 62 14:21:19 -1092.265055 0.174242
BFGS: 63 14:21:22 -1092.271638 0.126868
BFGS: 64 14:21:24 -1092.277221 0.126462
BFGS: 65 14:21:24 -1092.282878 0.127897
BFGS: 66 14:21:25 -1092.288371 0.129596
BFGS: 67 14:21:27 -1092.293223 0.166977
BFGS: 68 14:21:31 -1092.297711 0.135118
BFGS: 69 14:21:34 -1092.301968 0.139775
BFGS: 70 14:21:37 -1092.305902 0.164848
BFGS: 71 14:21:38 -1092.309897 0.189156
BFGS: 72 14:21:39 -1092.314215 0.181365
BFGS: 73 14:21:42 -1092.318594 0.156157
BFGS: 74 14:21:43 -1092.322559 0.126846
BFGS: 75 14:21:46 -1092.326041 0.088853
BFGS: 76 14:21:47 -1092.329305 0.121012
BFGS: 77 14:21:51 -1092.332259 0.112808
BFGS: 78 14:21:52 -1092.334626 0.078532
BFGS: 79 14:21:53 -1092.336384 0.079207
BFGS: 80 14:21:53 -1092.337807 0.059229
BFGS: 81 14:21:54 -1092.339134 0.063933
BFGS: 82 14:21:57 -1092.340470 0.084288
BFGS: 83 14:21:59 -1092.341827 0.105156
BFGS: 84 14:22:00 -1092.343214 0.104317
BFGS: 85 14:22:01 -1092.344594 0.091787
BFGS: 86 14:22:02 -1092.345942 0.076086
BFGS: 87 14:22:03 -1092.347222 0.069454
BFGS: 88 14:22:07 -1092.348397 0.053871
BFGS: 89 14:22:08 -1092.349443 0.038048
Energy of MOF + H2O: -1092.349 eV
We can now isolate the MOF atoms from the relaxed MOF + H2O geometry and see that the MOF has adopted a geometry that is less energetically favorable than the empty MOF by ~0.2 eV. The energy of the MOF in the adsorbed state corresponds to .
mof_adsorbed_state = mof_h2o[:-3]
mof_adsorbed_state.calc = calc
E_mof_adsorbed_state = mof_adsorbed_state.get_potential_energy()
print(f"Energy of MOF in the adsorbed state: {E_mof_adsorbed_state:.3f} eV")Energy of MOF in the adsorbed state: -1077.149 eV
H2O adsorption in this MOF appears to correspond to Case #2 as outlined above. We can now perform re-relaxation of the empty MOF starting from the geometry.
relax = BFGS(mof_adsorbed_state)
relax.run(fmax=0.05)
E_mof_rerelax = mof_adsorbed_state.get_potential_energy()
print(f"Energy of re-relaxed empty MOF: {E_mof_rerelax:.3f} eV") Step Time Energy fmax
BFGS: 0 14:22:08 -1077.149305 1.022068
BFGS: 1 14:22:09 -1077.190817 0.895214
BFGS: 2 14:22:11 -1077.241659 0.662570
BFGS: 3 14:22:11 -1077.289212 0.482407
BFGS: 4 14:22:14 -1077.306983 0.357667
BFGS: 5 14:22:15 -1077.323631 0.302100
BFGS: 6 14:22:16 -1077.337488 0.321930
BFGS: 7 14:22:16 -1077.349797 0.257280
BFGS: 8 14:22:17 -1077.356978 0.134732
BFGS: 9 14:22:17 -1077.361767 0.129605
BFGS: 10 14:22:18 -1077.366272 0.176027
BFGS: 11 14:22:18 -1077.371041 0.166113
BFGS: 12 14:22:19 -1077.375743 0.134315
BFGS: 13 14:22:20 -1077.380384 0.130561
BFGS: 14 14:22:21 -1077.384949 0.151711
BFGS: 15 14:22:21 -1077.388865 0.124835
BFGS: 16 14:22:22 -1077.391923 0.091722
BFGS: 17 14:22:22 -1077.394616 0.088809
BFGS: 18 14:22:23 -1077.397345 0.114583
BFGS: 19 14:22:24 -1077.400072 0.116739
BFGS: 20 14:22:25 -1077.402749 0.103742
BFGS: 21 14:22:27 -1077.405378 0.103394
BFGS: 22 14:22:28 -1077.407919 0.088820
BFGS: 23 14:22:30 -1077.410295 0.097398
BFGS: 24 14:22:32 -1077.412432 0.075307
BFGS: 25 14:22:33 -1077.414360 0.080454
BFGS: 26 14:22:33 -1077.416092 0.081145
BFGS: 27 14:22:33 -1077.417564 0.080486
BFGS: 28 14:22:34 -1077.418801 0.048850
Energy of re-relaxed empty MOF: -1077.419 eV
The MOF returns to its original empty reference energy upon re-relaxation, confirming that this deformation is physically relevant and is induced by the adsorbate molecule. In Case #3, this re-relaxed energy will be more negative (more favorable) than the original empty MOF relaxation. Thus, we take the reference empty MOF energy ( in Eqn. 1) to be the minimum of the original empty MOF energy and the re-relaxed MOf energy:
E_mof = min(E_mof_empty, E_mof_rerelax)
# get adsorbate reference energy
h2o = mof_h2o[-3:]
h2o.calc = calc
E_h2o = h2o.get_potential_energy()
# compute adsorption energy
E_ads = E_combo - E_mof - E_h2o
print(f"Adsorption energy of H2O in WOBHEB_0.11_0: {E_ads:.3f} eV")Adsorption energy of H2O in WOBHEB_0.11_0: -0.540 eV
This adsorption energy closely matches that from DFT (–0.699 eV) [1]. The strong adsorption energy is a consequence of both H2O chemisorption and MOF deformation. We can decompose the adsorption energy into contributions from these two factors. Assuming rigid H2O molecules, we define and , respectively, as
describes host host–guest interactions for the MOF in the adsorbed state only. quantifies the magnitude of deformation between the MOF in the adsorbed state and the most energetically favorable empty MOF geometry determined from the workflow presented here. It can be shown that
For H2O adsorption in WOBHEB_0.11, we have
E_int = E_combo - E_mof_adsorbed_state - E_h2o
print(f"E_int: {E_int}")E_int: -0.8269326882840939
E_mof_deform = E_mof_adsorbed_state - E_mof_empty
print(f"E_mof_deform: {E_mof_deform}")E_mof_deform: 0.286432945952356
E_ads = E_int + E_mof_deform
print(f"E_ads: {E_ads}")E_ads: -0.5404997423317379
is equivalent to when the MOF is assumed to be rigid. In this case, failure to consider adsorbate-induced deformation would result in an overestimation of the adsorption energy magnitude.
Acknowledgements & Authors¶
Logan Brabson and Sihoon Choi (Georgia Tech) and the OpenDAC project.
- Sriram, A., Choi, S., Yu, X., Brabson, L. M., Das, A., Ulissi, Z., Uyttendaele, M., Medford, A. J., & Sholl, D. S. (2024). The Open DAC 2023 Dataset and Challenges for Sorbent Discovery in Direct Air Capture. ACS Central Science, 10(5), 923–941. 10.1021/acscentsci.3c01629
- Queen, W. L., Hudson, M. R., Bloch, E. D., Mason, J. A., Gonzalez, M. I., Lee, J. S., Gygi, D., Howe, J. D., Lee, K., Darwish, T. A., James, M., Peterson, V. K., Teat, S. J., Smit, B., Neaton, J. B., Long, J. R., & Brown, C. M. (2014). Comprehensive study of carbon dioxide adsorption in the metal–organic frameworks M2(dobdc) (M = Mg, Mn, Fe, Co, Ni, Cu, Zn). Chem. Sci., 5(12), 4569–4581. 10.1039/c4sc02064b
- Yu, D., Yazaydin, A. O., Lane, J. R., Dietzel, P. D. C., & Snurr, R. Q. (2013). A combined experimental and quantum chemical study of CO2 adsorption in the metal–organic framework CPO-27 with different metals. Chemical Science, 4(9), 3544. 10.1039/c3sc51319j
- Alonso, G., Bahamon, D., Keshavarz, F., Giménez, X., Gamallo, P., & Sayós, R. (2018). Density Functional Theory-Based Adsorption Isotherms for Pure and Flue Gas Mixtures on Mg-MOF-74. Application in CO2 Capture Swing Adsorption Processes. The Journal of Physical Chemistry C, 122(7), 3945–3957. 10.1021/acs.jpcc.8b00938
- Witman, M., Ling, S., Jawahery, S., Boyd, P. G., Haranczyk, M., Slater, B., & Smit, B. (2017). The Influence of Intrinsic Framework Flexibility on Adsorption in Nanoporous Materials. Journal of the American Chemical Society, 139(15), 5547–5557. 10.1021/jacs.7b01688