This tutorial will walk you through a few examples of how you can use UMA. Each step is covered in more detail elsewhere in the documentation, but this is well suited to a ~1-2 hour tutorial session for researchers new to UMA but with some background in ASE and molecular simulations.
Before you start / installation¶
You need to get a HuggingFace account and request access to the UMA models.
You need a Huggingface account, request access to https://
Permissions: Read access to contents of all public gated repos you can access
Then, add the token as an environment variable (using huggingface-cli login:
# Enter token via huggingface-cli
! huggingface-cli loginor you can set the token via HF_TOKEN variable:
# Set token via env variable
import os
os.environ['HF_TOKEN'] = 'MYTOKEN'Installation process¶
It may be enough to use pip install fairchem-core. This gets you the latest version on PyPi (https://
Here we install some sub-packages. This can take 2-5 minutes to run.
! pip install fairchem-core fairchem-data-oc fairchem-applications-cattsunami x3dase# Check that packages are installed
!pip list | grep fairchemfairchem-applications-cattsunami 1.1.2.dev292+g268dd4092
fairchem-core 2.20.1.dev11+g268dd4092
fairchem-data-oc 1.0.3.dev292+g268dd4092
fairchem-data-omat 0.2.1.dev197+g268dd4092
import fairchem.core
fairchem.core.__version__'2.20.1.dev11+g268dd4092'Illustrative examples¶
These should just run, and are here to show some basic uses.
Spin gap energy - OMOL¶
This is the difference in energy between a triplet and single ground state for a CH2 radical. This downloads a ~1GB checkpoint the first time you run it.
We don’t set a device here, so we get a warning about using a CPU device. You can ignore that. If a CUDA environment is available, a GPU may be used to speed up the calculations.
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
from ase.build import molecule
# singlet CH2
singlet = molecule("CH2_s1A1d")
singlet.info.update({"spin": 1, "charge": 0})
singlet.calc = FAIRChemCalculator(predictor, task_name="omol")
# triplet CH2
triplet = molecule("CH2_s3B1d")
triplet.info.update({"spin": 3, "charge": 0})
triplet.calc = FAIRChemCalculator(predictor, task_name="omol")
print(triplet.get_potential_energy() - singlet.get_potential_energy())-0.6104808867351039
Example of adsorbate relaxation - OC20¶
Here we just setup a Cu(100) slab with a CO on it and relax it.
We specify an explicit device in the predictor here, and avoid the warning.
from ase.build import add_adsorbate, fcc100, molecule
from ase.optimize import LBFGS
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="oc20")
# Set up your system as an ASE atoms object
slab = fcc100("Cu", (3, 3, 3), vacuum=8, periodic=True)
adsorbate = molecule("CO")
add_adsorbate(slab, adsorbate, 2.0, "bridge")
slab.calc = calc
# Set up LBFGS dynamics object
opt = LBFGS(slab)
opt.run(0.05, 100)
print(slab.get_potential_energy())WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
Step Time Energy fmax
LBFGS: 0 13:47:41 -89.544140 11.402925
LBFGS: 1 13:47:41 -92.457702 6.589585
LBFGS: 2 13:47:41 -92.585267 7.565847
LBFGS: 3 13:47:41 -92.966559 3.751868
LBFGS: 4 13:47:41 -93.124911 3.558189
LBFGS: 5 13:47:41 -93.231281 2.245371
LBFGS: 6 13:47:41 -93.471486 1.133989
LBFGS: 7 13:47:41 -93.563000 0.996113
LBFGS: 8 13:47:41 -93.673182 0.698816
LBFGS: 9 13:47:41 -93.760220 0.492676
LBFGS: 10 13:47:42 -93.806518 0.365447
LBFGS: 11 13:47:42 -93.825450 0.344569
LBFGS: 12 13:47:42 -93.849868 0.486299
LBFGS: 13 13:47:42 -93.868569 0.424331
LBFGS: 14 13:47:42 -93.878644 0.154896
LBFGS: 15 13:47:42 -93.884595 0.169650
LBFGS: 16 13:47:42 -93.891117 0.207732
LBFGS: 17 13:47:42 -93.897925 0.250701
LBFGS: 18 13:47:42 -93.903971 0.173538
LBFGS: 19 13:47:42 -93.906880 0.053488
LBFGS: 20 13:47:43 -93.907326 0.042109
-93.9073264872675
Example bulk relaxation - OMAT¶
from ase.build import bulk
from ase.filters import FrechetCellFilter
from ase.optimize import FIRE
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="omat")
atoms = bulk("Fe")
atoms.calc = calc
opt = FIRE(FrechetCellFilter(atoms))
opt.run(0.05, 100)
print(atoms.get_stress()) # !!!! We get stress now!WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
Step Time Energy fmax
FIRE: 0 13:47:45 -8.263805 0.571327
FIRE: 1 13:47:46 -8.270171 0.139049
FIRE: 2 13:47:46 -8.259920 1.610376
FIRE: 3 13:47:46 -8.269784 0.262882
FIRE: 4 13:47:46 -8.269145 0.282860
FIRE: 5 13:47:46 -8.269291 0.268272
FIRE: 6 13:47:46 -8.269553 0.239073
FIRE: 7 13:47:46 -8.269875 0.195151
FIRE: 8 13:47:47 -8.270182 0.136503
FIRE: 9 13:47:47 -8.270394 0.063354
FIRE: 10 13:47:47 -8.270440 0.023303
[-2.01406764e-03 -2.01445624e-03 -2.01398653e-03 2.12174828e-09
-6.56061425e-09 6.22314134e-09]
Molecular dynamics - OMOL¶
import matplotlib.pyplot as plt
from ase import units
from ase.build import molecule
from ase.io import Trajectory
from ase.md.langevin import Langevin
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="omol")
atoms = molecule("H2O")
atoms.info.update(charge=0, spin=1) # For omol
atoms.calc = calc
dyn = Langevin(
atoms,
timestep=0.1 * units.fs,
temperature_K=400,
friction=0.001 / units.fs,
)
trajectory = Trajectory("my_md.traj", "w", atoms)
dyn.attach(trajectory.write, interval=1)
dyn.run(steps=50)
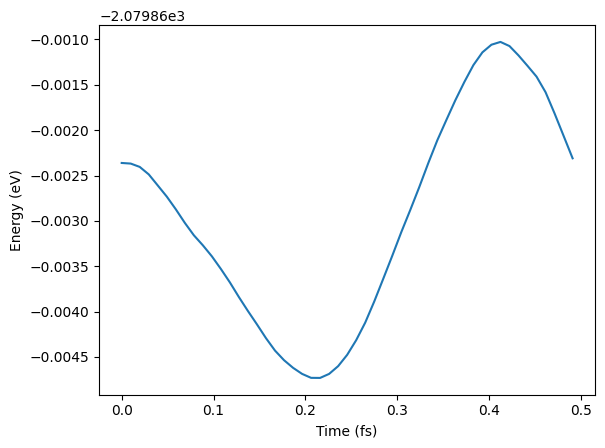
# See some results - not paper ready!
traj = Trajectory("my_md.traj")
plt.plot(
[i * 0.1 * units.fs for i in range(len(traj))],
[a.get_potential_energy() for a in traj],
)
plt.xlabel("Time (fs)")
plt.ylabel("Energy (eV)");WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
/home/runner/work/_tool/Python/3.12.13/x64/lib/python3.12/site-packages/ase/md/langevin.py:110: FutureWarning: The implementation of `fixcm=True` in `Langevin` does not strictly sample the correct NVT distributions. The deviations are typically small for large systems but can be more pronounced for small systems. Use `fixcm=False` together with `ase.constraints.FixCom`. `fixcm` is deprecated since ASE 3.28.0 and will be removed in a future release.
warnings.warn(msg, FutureWarning)
Catalyst Adsorption energies¶
The basic approach in computing an adsorption energy is to compute this energy difference:
dH = E_adslab - E_slab - E_adsWe use UMA for two of these energies E_adslab and E_slab. For E_ads We have to do something a little different. The OC20 task is not trained for molecules or molecular fragments. We use atomic energy reference energies instead. These are tabulated below.
The OC20 reference scheme is this reaction:
x CO + (x + y/2 - z) H2 + (z-x) H2O + w/2 N2 + * -> CxHyOzNw* For this example we have
-H2 + H2O + * -> O*. "O": -7.204 eVWhere "O": -7.204 is a constant.
To get the desired reaction energy we want we add the formation energy of water. We use either DFT or experimental values for this reaction energy.
1/2O2 + H2 -> H2OAlternatives to this approach are using DFT to estimate the energy of 1/2 O2, just make sure to use consistent settings with your task. You should not use OMOL for this.
from ase.build import add_adsorbate, fcc111
from ase.optimize import BFGS
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="oc20")WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
# reference energies from a linear combination of H2O/N2/CO/H2!
atomic_reference_energies = {
"H": -3.477,
"N": -8.083,
"O": -7.204,
"C": -7.282,
}
re1 = -3.03 # Water formation energy from experiment
slab = fcc111("Pt", size=(2, 2, 5), vacuum=20.0)
slab.pbc = True
adslab = slab.copy()
add_adsorbate(adslab, "O", height=1.2, position="fcc")
slab.calc = calc
opt = BFGS(slab)
print("Relaxing slab")
opt.run(fmax=0.05, steps=100)
slab_e = slab.get_potential_energy()
adslab.calc = calc
opt = BFGS(adslab)
print("\nRelaxing adslab")
opt.run(fmax=0.05, steps=100)
adslab_e = adslab.get_potential_energy()Relaxing slab
Step Time Energy fmax
BFGS: 0 13:47:57 -104.695195 0.709591
BFGS: 1 13:47:58 -104.753187 0.607748
BFGS: 2 13:47:58 -104.906055 0.369389
BFGS: 3 13:47:58 -104.938753 0.439758
BFGS: 4 13:47:58 -105.016145 0.464012
BFGS: 5 13:47:58 -105.076562 0.356059
BFGS: 6 13:47:58 -105.112621 0.189428
BFGS: 7 13:47:58 -105.126758 0.045369
Relaxing adslab
Step Time Energy fmax
BFGS: 0 13:47:58 -110.048788 1.756661
BFGS: 1 13:47:58 -110.230718 0.986316
BFGS: 2 13:47:59 -110.379375 0.731556
BFGS: 3 13:47:59 -110.430298 0.807226
BFGS: 4 13:47:59 -110.543733 0.691549
BFGS: 5 13:47:59 -110.617505 0.492728
BFGS: 6 13:47:59 -110.673478 0.667090
BFGS: 7 13:47:59 -110.721709 0.710360
BFGS: 8 13:48:00 -110.756914 0.443592
BFGS: 9 13:48:00 -110.769298 0.214401
BFGS: 10 13:48:00 -110.772451 0.091648
BFGS: 11 13:48:00 -110.772963 0.062454
BFGS: 12 13:48:00 -110.773258 0.046192
Now we compute the adsorption energy.
# Energy for ((H2O-H2) + * -> *O) + (H2 + 1/2O2 -> H2O) leads to 1/2O2 + * -> *O!
adslab_e - slab_e - atomic_reference_energies["O"] + re1-1.4724996175374723How did we do? We need a reference point. In the paper below, there is an atomic adsorption energy for O on Pt(111) of about -4.264 eV. This is for the reaction O + * -> O*. To convert this to the dissociative adsorption energy, we have to add the reaction:
1/2 O2 -> O D = 2.58 eV (expt)to get a comparable energy of about -1.68 eV. There is about ~0.2 eV difference (we predicted -1.47 eV above, and the reference comparison is -1.68 eV) to account for. The biggest difference is likely due to the differences in exchange-correlation functional. The reference data used the PBE functional, and eSCN was trained on RPBE data. To additional places where there are differences include:
Difference in lattice constant
The reference energy used for the experiment references. These can differ by up to 0.5 eV from comparable DFT calculations.
How many layers are relaxed in the calculation
Some of these differences tend to be systematic, and you can calibrate and correct these, especially if you can augment these with your own DFT calculations.
It is always a good idea to visualize the geometries to make sure they look reasonable.
import matplotlib.pyplot as plt
from ase.visualize.plot import plot_atoms
fig, axs = plt.subplots(1, 2)
plot_atoms(slab, axs[0])
plot_atoms(slab, axs[1], rotation=("-90x"))
axs[0].set_axis_off()
axs[1].set_axis_off()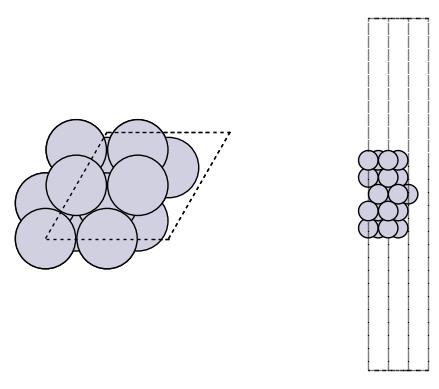
fig, axs = plt.subplots(1, 2)
plot_atoms(adslab, axs[0])
plot_atoms(adslab, axs[1], rotation=("-90x"))
axs[0].set_axis_off()
axs[1].set_axis_off()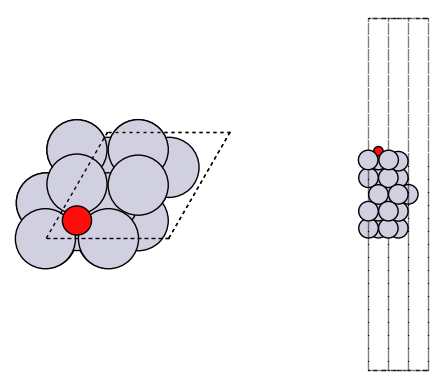
Molecular vibrations¶
from ase import Atoms
from ase.optimize import BFGS
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="omol")
from ase.vibrations import Vibrations
n2 = Atoms("N2", [(0, 0, 0), (0, 0, 1.1)])
n2.info.update({"spin": 1, "charge": 0})
n2.calc = calc
BFGS(n2).run(fmax=0.01)WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
Step Time Energy fmax
BFGS: 0 13:48:07 -2981.069021 1.646100
BFGS: 1 13:48:07 -2980.962521 6.604125
BFGS: 2 13:48:07 -2981.077760 0.200688
BFGS: 3 13:48:07 -2981.077885 0.023328
BFGS: 4 13:48:07 -2981.077887 0.000100
np.True_vib = Vibrations(n2)
vib.run()
vib.summary()---------------------
# meV cm^-1
---------------------
0 0.0i 0.0i
1 0.0i 0.0i
2 0.0 0.0
3 2.0 16.1
4 2.0 16.1
5 309.6 2496.7
---------------------
Zero-point energy: 0.157 eV
Bulk alloy phase behavior¶
Adapted from https://
We manually compute the formation energy of pure compounds and some alloy compositions to assess stability.
from ase.atoms import Atom, Atoms
from ase.filters import FrechetCellFilter
from ase.optimize import FIRE
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
cu = Atoms(
[Atom("Cu", [0.000, 0.000, 0.000])],
cell=[[1.818, 0.000, 1.818], [1.818, 1.818, 0.000], [0.000, 1.818, 1.818]],
pbc=True,
)
cu.calc = FAIRChemCalculator(predictor, task_name="omat")
opt = FIRE(FrechetCellFilter(cu))
opt.run(0.05, 100)
cu.get_potential_energy()WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
Step Time Energy fmax
FIRE: 0 13:48:11 -3.748689 0.185703
FIRE: 1 13:48:11 -3.749558 0.126065
FIRE: 2 13:48:11 -3.750259 0.023407
-3.750259408643074pd = Atoms(
[Atom("Pd", [0.000, 0.000, 0.000])],
cell=[[1.978, 0.000, 1.978], [1.978, 1.978, 0.000], [0.000, 1.978, 1.978]],
pbc=True,
)
pd.calc = FAIRChemCalculator(predictor, task_name="omat")
opt = FIRE(FrechetCellFilter(pd))
opt.run(0.05, 100)
pd.get_potential_energy() Step Time Energy fmax
FIRE: 0 13:48:11 -5.208746 0.206138
FIRE: 1 13:48:11 -5.209731 0.111874
FIRE: 2 13:48:11 -5.210079 0.040758
-5.210078651522462Alloy formation energies¶
cupd1 = Atoms(
[Atom("Cu", [0.000, 0.000, 0.000]), Atom("Pd", [-1.652, 0.000, 2.039])],
cell=[[0.000, -2.039, 2.039], [0.000, 2.039, 2.039], [-3.303, 0.000, 0.000]],
pbc=True,
) # Note pbc=True is important, it is not the default and OMAT
cupd1.calc = FAIRChemCalculator(predictor, task_name="omat")
opt = FIRE(FrechetCellFilter(cupd1))
opt.run(0.05, 100)
cupd1.get_potential_energy() Step Time Energy fmax
FIRE: 0 13:48:13 -9.200180 0.149828
FIRE: 1 13:48:13 -9.200405 0.137151
FIRE: 2 13:48:13 -9.200767 0.114193
FIRE: 3 13:48:13 -9.201159 0.085833
FIRE: 4 13:48:13 -9.201547 0.064594
FIRE: 5 13:48:14 -9.201993 0.080829
FIRE: 6 13:48:14 -9.202580 0.083421
FIRE: 7 13:48:14 -9.203317 0.072357
FIRE: 8 13:48:15 -9.204220 0.065461
FIRE: 9 13:48:15 -9.205216 0.090108
FIRE: 10 13:48:15 -9.206354 0.104151
FIRE: 11 13:48:15 -9.207779 0.094694
FIRE: 12 13:48:16 -9.209430 0.060871
FIRE: 13 13:48:16 -9.211121 0.055557
FIRE: 14 13:48:16 -9.212895 0.038686
-9.212895336664005cupd2 = Atoms(
[
Atom("Cu", [-0.049, 0.049, 0.049]),
Atom("Cu", [-11.170, 11.170, 11.170]),
Atom("Pd", [-7.415, 7.415, 7.415]),
Atom("Pd", [-3.804, 3.804, 3.804]),
],
cell=[[-5.629, 3.701, 5.629], [-3.701, 5.629, 5.629], [-5.629, 5.629, 3.701]],
pbc=True,
)
cupd2.calc = FAIRChemCalculator(predictor, task_name="omat")
opt = FIRE(FrechetCellFilter(cupd2))
opt.run(0.05, 100)
cupd2.get_potential_energy() Step Time Energy fmax
FIRE: 0 13:48:17 -18.132613 0.170385
FIRE: 1 13:48:17 -18.133448 0.153581
FIRE: 2 13:48:17 -18.134793 0.121402
FIRE: 3 13:48:17 -18.136120 0.076687
FIRE: 4 13:48:18 -18.136904 0.023653
-18.136903551070727# Delta Hf cupd-1 = -0.11 eV/atom
hf1 = (
cupd1.get_potential_energy() - cu.get_potential_energy() - pd.get_potential_energy()
)
hf1-0.25255727649846893# DFT: Delta Hf cupd-2 = -0.04 eV/atom
hf2 = (
cupd2.get_potential_energy()
- 2 * cu.get_potential_energy()
- 2 * pd.get_potential_energy()
)
hf2-0.21622743073965545hf1 - hf2, (-0.11 - -0.04)(-0.036329845758813484, -0.07)These indicate that cupd-1 and cupd-2 are both more stable than phase separated Cu and Pd, and that cupd-1 is more stable than cupd-2. The absolute formation energies differ from the DFT references, but the relative differences are quite close. The absolute differences could be due to DFT parameter choices (XC, psp, etc.).
Phonon calculation¶
This takes 4-10 minutes. Adapted from https://
Phonons have applications in computing the stability and free energy of solids. See:
https://
www .sciencedirect .com /science /article /pii /S1359646215003127 https://
iopscience .iop .org /book /mono /978 -0 -7503 -2572 -1 /chapter /bk978 -0 -7503 -2572 -1ch1
from ase.build import bulk
from ase.phonons import Phonons
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="omat")
# Setup crystal
atoms = bulk("Al", "fcc", a=4.05)
# Phonon calculator
N = 7
ph = Phonons(atoms, calc, supercell=(N, N, N), delta=0.05)
ph.run()
# Read forces and assemble the dynamical matrix
ph.read(acoustic=True)
ph.clean()
path = atoms.cell.bandpath("GXULGK", npoints=100)
bs = ph.get_band_structure(path)
dos = ph.get_dos(kpts=(20, 20, 20)).sample_grid(npts=100, width=1e-3)WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
WARNING, 1 imaginary frequencies at q = ( 0.02, 0.02, 0.02) ; (omega_q = 1.434e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.01, 0.01, 0.03) ; (omega_q = 1.660e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.03, 0.03, 0.05) ; (omega_q = 2.249e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.04, 0.04, 0.08) ; (omega_q = 2.055e-02*i)
WARNING, 1 imaginary frequencies at q = (-0.08, -0.03, -0.03) ; (omega_q = 1.617e-02*i)
WARNING, 1 imaginary frequencies at q = (-0.03, -0.08, -0.03) ; (omega_q = 1.618e-02*i)
WARNING, 1 imaginary frequencies at q = (-0.03, -0.03, -0.08) ; (omega_q = 1.618e-02*i)
WARNING, 1 imaginary frequencies at q = (-0.03, -0.03, -0.03) ; (omega_q = 1.446e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.03, 0.03, 0.03) ; (omega_q = 1.446e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.03, 0.03, 0.07) ; (omega_q = 1.618e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.03, 0.07, 0.03) ; (omega_q = 1.618e-02*i)
WARNING, 1 imaginary frequencies at q = ( 0.07, 0.03, 0.03) ; (omega_q = 1.617e-02*i)
# Plot the band structure and DOS:
import matplotlib.pyplot as plt # noqa
fig = plt.figure(figsize=(7, 4))
ax = fig.add_axes([0.12, 0.07, 0.67, 0.85])
emax = 0.04
bs.plot(ax=ax, emin=0.0, emax=emax)
dosax = fig.add_axes([0.8, 0.07, 0.17, 0.85])
dosax.fill_between(
dos.get_weights(),
dos.get_energies(),
y2=0,
color="grey",
edgecolor="k",
lw=1,
)
dosax.set_ylim(0, emax)
dosax.set_yticks([])
dosax.set_xticks([])
dosax.set_xlabel("DOS", fontsize=18);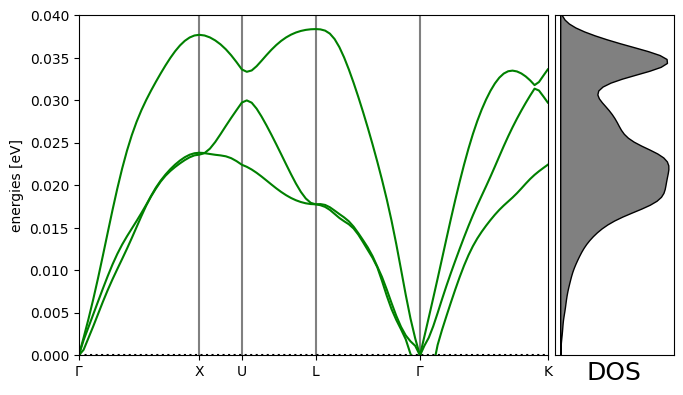
Transition States (NEBs)¶
Nudged elastic band calculations are among the most costly calculations we do. UMA makes these quicker!
We explore diffusion of an O adatom from an hcp to an fcc site on Pt(111).
Initial state¶
from ase.build import add_adsorbate, fcc111, molecule
from ase.optimize import LBFGS
from fairchem.core import FAIRChemCalculator, pretrained_mlip
predictor = pretrained_mlip.get_predict_unit("uma-s-1p1")
calc = FAIRChemCalculator(predictor, task_name="oc20")
# Set up your system as an ASE atoms object
initial = fcc111("Pt", (3, 3, 3), vacuum=8, periodic=True)
adsorbate = molecule("O")
add_adsorbate(initial, adsorbate, 2.0, "fcc")
initial.calc = calc
# Set up LBFGS dynamics object
opt = LBFGS(initial)
opt.run(0.05, 100)
print(initial.get_potential_energy())WARNING:root:device was not explicitly set, using device='cuda'.
WARNING:root:If 'dataset_list' is provided in the config, the code assumes that each dataset maps to itself. Please use 'dataset_mapping' as 'dataset_list' is deprecated and will be removed in the future.
Step Time Energy fmax
LBFGS: 0 13:48:32 -141.309107 3.517761
LBFGS: 1 13:48:32 -141.702377 3.524942
LBFGS: 2 13:48:32 -142.967948 2.974642
LBFGS: 3 13:48:32 -143.672289 0.963273
LBFGS: 4 13:48:32 -143.776589 1.269393
LBFGS: 5 13:48:32 -143.848510 0.873111
LBFGS: 6 13:48:32 -143.923350 0.164694
LBFGS: 7 13:48:33 -143.926599 0.153173
LBFGS: 8 13:48:33 -143.933660 0.126651
LBFGS: 9 13:48:33 -143.937971 0.116467
LBFGS: 10 13:48:33 -143.941406 0.073791
LBFGS: 11 13:48:34 -143.942884 0.083219
LBFGS: 12 13:48:34 -143.944363 0.084958
LBFGS: 13 13:48:34 -143.945986 0.065906
LBFGS: 14 13:48:34 -143.947506 0.037419
-143.94750638511687
Final state¶
# Set up your system as an ASE atoms object
final = fcc111("Pt", (3, 3, 3), vacuum=8, periodic=True)
adsorbate = molecule("O")
add_adsorbate(final, adsorbate, 2.0, "hcp")
final.calc = FAIRChemCalculator(predictor, task_name="oc20")
# Set up LBFGS dynamics object
opt = LBFGS(final)
opt.run(0.05, 100)
print(final.get_potential_energy()) Step Time Energy fmax
LBFGS: 0 13:48:35 -141.268799 3.362366
LBFGS: 1 13:48:35 -141.650619 3.342133
LBFGS: 2 13:48:36 -142.883885 2.576832
LBFGS: 3 13:48:36 -143.406005 1.208928
LBFGS: 4 13:48:36 -143.469218 0.959532
LBFGS: 5 13:48:36 -143.589295 0.127616
LBFGS: 6 13:48:37 -143.593123 0.112180
LBFGS: 7 13:48:37 -143.595710 0.091209
LBFGS: 8 13:48:37 -143.597250 0.071142
LBFGS: 9 13:48:37 -143.598559 0.047105
-143.5985592556432
Setup and relax the band¶
from ase.mep import NEB
images = [initial]
for i in range(3):
image = initial.copy()
image.calc = FAIRChemCalculator(predictor, task_name="oc20")
images.append(image)
images.append(final)
neb = NEB(images)
neb.interpolate()
opt = LBFGS(neb, trajectory="neb.traj")
opt.run(0.05, 100)/home/runner/work/_tool/Python/3.12.13/x64/lib/python3.12/site-packages/ase/mep/neb.py:329: UserWarning: The default method has changed from 'aseneb' to 'improvedtangent'. The 'aseneb' method is an unpublished, custom implementation that is not recommended as it frequently results in very poor bands. Please explicitly set method='improvedtangent' to silence this warning, or set method='aseneb' if you strictly require the old behavior (results may vary). See: https://gitlab.com/ase/ase/-/merge_requests/3952
warnings.warn(
Step Time Energy fmax
LBFGS: 0 13:48:38 -143.171394 3.064800
LBFGS: 1 13:48:38 -143.340627 1.486512
LBFGS: 2 13:48:39 -143.393668 0.434151
LBFGS: 3 13:48:40 -143.405931 0.453339
LBFGS: 4 13:48:41 -143.426328 0.462192
LBFGS: 5 13:48:41 -143.442346 0.346495
LBFGS: 6 13:48:42 -143.452654 0.180222
LBFGS: 7 13:48:42 -143.457226 0.151730
LBFGS: 8 13:48:43 -143.459749 0.175266
LBFGS: 9 13:48:43 -143.461192 0.188564
LBFGS: 10 13:48:44 -143.462572 0.176440
LBFGS: 11 13:48:44 -143.463358 0.094809
LBFGS: 12 13:48:45 -143.463785 0.090830
LBFGS: 13 13:48:45 -143.464270 0.101375
LBFGS: 14 13:48:45 -143.464864 0.128213
LBFGS: 15 13:48:45 -143.465403 0.087877
LBFGS: 16 13:48:46 -143.465721 0.047130
np.True_from ase.mep import NEBTools
NEBTools(neb.images).plot_band();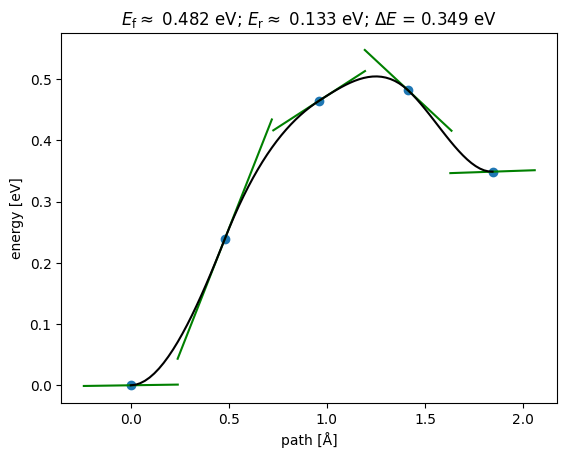
This could be a good initial guess to initialize an NEB in DFT.
Ideas for things you can do with UMA¶
Advanced applications¶
These take a while to run.
AdsorbML¶
It is so cheap to run these calculations that we can screen a broad range of adsorbate sites and rank them in stability. The AdsorbML approach automates this. This takes quite a while to run here, and we don’t do it in the workshop.
Expert adsorption energies¶
This tutorial reproduces Fig 6b from the following paper: Zhou, Jing, et al. “Enhanced Catalytic Activity of Bimetallic Ordered Catalysts for Nitrogen Reduction Reaction by Perturbation of Scaling Relations.” ACS Catalysis 134 (2023): 2190-2201 (Zhou et al. (2023)).
This takes up to an hour with a GPU, and much longer with a CPU.
CatTsunami¶
The CatTsunami tutorial is an example of enumerating initial and final states, and computing reaction paths between them with UMA.
Acknowledgments¶
This tutorial was originally compiled by John Kitchin (CMU) for the NAM29 catalysis tutorial session, using a variety of resources from the FAIR chemistry repository.
- Musielewicz, J., Wang, X., Tian, T., & Ulissi, Z. (2022). FINETUNA: fine-tuning accelerated molecular simulations. Machine Learning: Science and Technology, 3(3), 03LT01. 10.1088/2632-2153/ac8fe0
- Wang, X., Musielewicz, J., Tran, R., Kumar Ethirajan, S., Fu, X., Mera, H., Kitchin, J. R., Kurchin, R. C., & Ulissi, Z. W. (2024). Generalization of graph-based active learning relaxation strategies across materials. Machine Learning: Science and Technology, 5(2), 025018. 10.1088/2632-2153/ad37f0
- Wander, B., Musielewicz, J., Cheula, R., & Kitchin, J. R. (2025). Accessing Numerical Energy Hessians with Graph Neural Network Potentials and Their Application in Heterogeneous Catalysis. The Journal of Physical Chemistry C, 129(7), 3510–3521. 10.1021/acs.jpcc.4c07477
- Zhou, J., Chen, X., Guo, M., Hu, W., Huang, B., & Yuan, D. (2023). Enhanced Catalytic Activity of Bimetallic Ordered Catalysts for Nitrogen Reduction Reaction by Perturbation of Scaling Relations. ACS Catalysis, 13(4), 2190–2201. 10.1021/acscatal.2c05877